
FASTX implements its parser through a high-level description of the FASTQ format thanks to Automa and lets the machine figure out most of the gritty details. 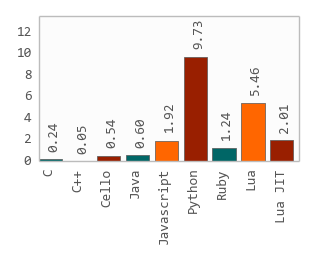
Second, I think it’s very instructive to compare the approach to FASTQ parsing represented by klib.h vs FASTX.jl. It’s hard to think of a real-life application where any useful work could be done so quick that these timings matter much. On my laptop, Julia crunches 3.6 million reads/sec uncompressed and something like 1.6 m/s compressed. W.r.t the broader implicaitons of Julia, first notice the absolute timings. Nonetheless, I do think it’s nice to have the foundation of Julia bioinfo packages be so fast that people would not be turned away from implementing highly performance-sensitive tools it in, like a short-read aligner or an assembler. Indeed - that becomes a much more involved discussion than pure performance and can’t easily be resolved with tables and measurements. I don’t know if we can find a faster implementation of zlib to use in CodecZlib - presumably MacOS can make certain assumptions about what OS and CPU their users have which Julia can’t. The remaining difference in performance is therefore due to an upstream ineffiency. That puts us way behind C speed, just a tad before Python.Įdit2: It seems the zlib.h source code that CodecZlib obtains from is not as optimized as the zlib library that ships with MacOS. Except for the boundscheck issue and the zlib issue, the only place left I can find is to optimize Automa.jl directly - which would always be welcome, but I don’t know how.Įdit: Oh yes, and the elephant in the room: The Julia code goes through the 5 million FASTQ read in 5 seconds, whereas it takes around 11 when called from command line due to compile time latency. The implementation of the FASTQ format and the parsing is very efficient. If anyone can fix the CodecZlib issue, that would be great. After this, Julia is about 1.4 slower than C on gzipped data - whereas we ought to be very near C speed here, something like a factor of 1.1. I created this issue - Jaakko Ruohio on Slack discovered about half the problem was that zlib was not compiled with -O3, but there are still more gains to be had, I just don’t know where to find them. Profiling confirms that almost all time is spent in the ccall line. That’s very strange, considering it just calls zlib directly. Not bad! We probably shouldn’t actually remove the boundschecks, because IMO the extra safety is worth a little extra time, especially considering that FASTQ files are essentially always gzipped.įor the gzipped data, it appears that CodecZlib is very slow, something like 4x slower than zlib. With this optimization, the running speed of Julia is about 1.25-1.3 times as long as his C code. Since it reads the input byte by byte, this has a rather large effect, taking another 30% time off the non-zipped input. 
There’s some type instability, but union splitting takes care of that, and it’s no performance concern.įor the FASTX code, I noticed the FASTQ parser does not use inbounds. Looking at his Julia code, it looks great. It might be a matter of hardware, or the fact that I ran FASTX v 1.1, not 1.0. I don’t see the same relative numbers b/w C, Python and Julia that he does - for me, Julia does about 30% better on non-zipped data. It’s much more interesting to look at the FASTQ benchmark using FASTX.jl from BioJulia. I don’t think it’s useful to pick that code apart, it’s just one random guy’s Julia code. The library Klib.jl is a bit obscure, seemingly just used by one guy, and not very well maintained. I recreated the C, Julia and Python part of the first benchmark.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
